Radiological Society of North America (RSNA) has reported that researchers have used radiomic feature maps to identify the cancer patients who respond to chemotherapy.
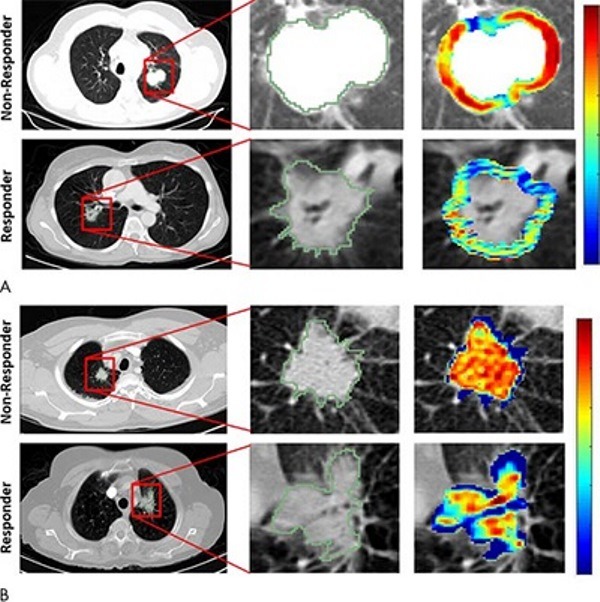
Image: Radiomic feature maps showing the differences in lesions of responders and non-responders. Photo: Courtesy of Radiological Society of North America.
According to RSNA, researchers stated that radiomic feature maps show the differences between the lesions in responders and non-responders in the pretreatment CT scans. They found that radiomic features (A, peritumoral Gabor and, B, intratumoral Laws features) overexpress in the nonresponder lesion compared to the lesions of responders.
The distribution of features shows significant differences between the two lesion categories and they are captured by the skewness and kurtosis statistics of the feature distribution.
By using data from CT images, researchers could be able to predict which lung cancer patients will respond to chemotherapy.
The first-line of treatment for advanced-stage non-small cell lung cancer (NSCLC) is platinum-based chemotherapy. But, only about one in four patients are expected to respond well to this treatment. Presently, there is no way of predicting which patients will benefit from this therapy.
CT scans are routinely used to find tumor staging and for monitoring treatment response. By using radiomics, researchers can quantitatively know the disease characteristics that not visible in the images.
Case Western Reserve University School of Engineering Department of Biomedical Engineering Mohammadhadi Khorrami and Monica Khunger, Cleveland Clinic Department of Internal Medicine said: “Our aim in this study was to determine whether an early prediction of response to chemotherapy is possible by using computer-extracted measurements of patterns both within and outside the lung nodule, along with the shape of the nodule, on baseline CT scans.”
The RSNA research aimed at identifying the role of radiomic texture features within and around the lung tumor, which could help in predicting the time of progression and overall survival and response to chemotherapy in patients with NSCLC.
The duo analyzed data from 125 patients who were treated with pemetrexed-based platinum doublet chemotherapy at Cleveland Clinic. The patients were divided randomly into two sets with equal number responders and non-responders in the training set. The training set included 53 patients with NSCLC, and the validation set included 72 patients.
Data from the patients was analyzed by a computer to identify the unique patterns of heterogeneity both inside and outside the tumor. Those patients were then compared between CT scans of patients who did and did not respond to chemotherapy. These patterns were then used to train a machine learning program to identify the chance of a lung cancer patient would respond to chemotherapy.
Khunger said: “This is the first study to demonstrate that computer-extracted patterns of heterogeneity, or diversity, from outside the tumor were predictive of response to chemotherapy.
“This is very critical because it could allow for predicting in advance of therapy which patients with lung cancer are likely to respond or not. This, in turn, could help identify patients who are likely to not respond to chemotherapy for alternative therapies such as radiation or immunotherapy.”
Khorrami continued: “When we looked at patterns inside the tumor, we got an accuracy of 0.68. But when we looked inside and outside, the accuracy went up to 0.77.”
The results showed that radiomic features derived from within the tumor and the area around the tumor were able to distinguish patients who responded to the therapy from those who did not and also predicted the time of profession and overall survival.
